Motivation
Electron cryo-microscopy (cryo-EM) is an important and rapidly developing field of structural biology. Thanks to numerous advancements in instrumentation and data processing software, the number of structures determined by cryo-EM is growing rapidly and the resolution of cryo-EM maps is improving dramatically. Computational tools for the analysis of 3D reconstructions and atomic model refinement are also undergoing rapid development. It is likely that current refinement software can provide improved models compared to those determined previously. This site makes available updated versions of existing cryo-EM models automatically obtained using the latest version of phenix.real_space_refine.
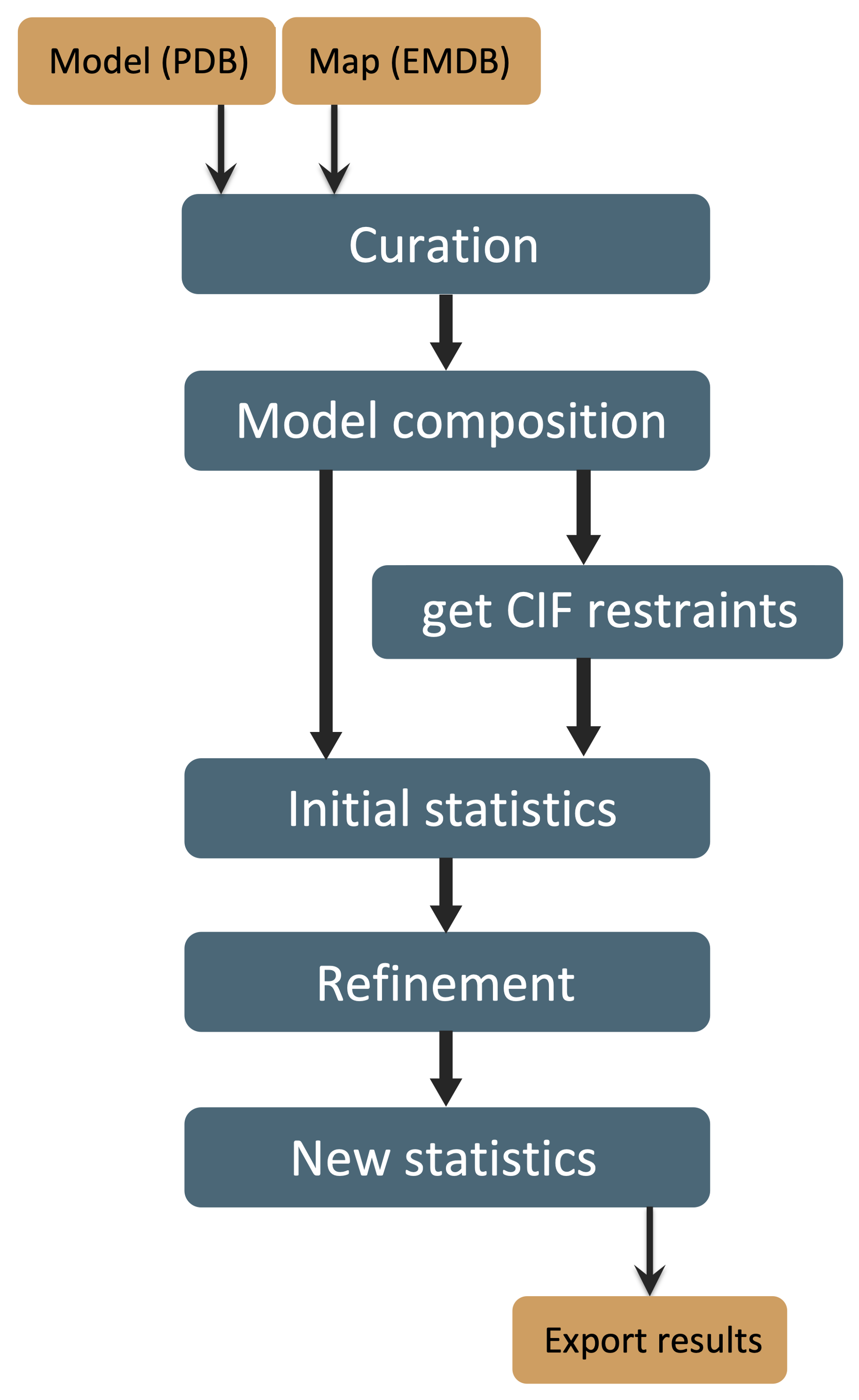
Procedure
Hover over the boxes in the figure to see a more detailed description in the shaded frame.
Literature
List of publications for tools that were used to perform the CERES computations:
-
CERES webserver:
Liebschner, D.; Afonine, P. V.; Moriarty, N. W.; Poon, B. K.; Chen, V. B.; Adams, P. D. CERES : A Cryo-EM Re-Refinement System for Continuous Improvement of Deposited Models. Acta Cryst. D 2021, 77 (1), 48–61 https://doi.org/10.1107/S2059798320015879 -
Program to perform refinements in real-space: phenix.real_space_refine
Afonine, P. V.; Poon, B. K.; Read, R. J.; Sobolev, O. V.; Terwilliger, T. C.; Urzhumtsev, A.; Adams, P. D. Real-Space Refinement in PHENIX for Cryo-EM and Crystallography. Acta Cryst. D 2018, 74 (6), 531–544. https://doi.org/10.1107/S2059798318006551
Documentation for phenix.real_space_refine -
Program to generate ligand restraints: eLBOW
Moriarty, N. W.; Grosse-Kunstleve, R. W.; Adams, P. D. Electronic Ligand Builder and Optimization Workbench ( ELBOW ): A Tool for Ligand Coordinate and Restraint Generation. Acta Cryst. D 2009, 65 (10)1074–1080 https://doi.org/10.1107/S0907444909029436
Documentation for phenix.elbow -
Metrics for the validation of cryo-EM maps and models
Afonine, P. V.; Klaholz, B. P.; Moriarty, N. W.; Poon, B. K.; Sobolev, O. V.; Terwilliger, T. C.; Adams, P. D.; Urzhumtsev, A. New Tools for the Analysis and Validation of Cryo-EM Maps and Atomic Models. Acta Cryst. D 2018, 74 (9), 814–840. https://doi.org/10.1107/S2059798318009324