Fitting a loop
Learn how to fill in a gap in a model with the fit_loop tool.
Setting up example data
First, let's set up the example data (described in more detail here):
Get the files from the Phenix regression directory and change into the new folder:
phenix.setup_tutorial tutorial_name=model-building-scripting
cd model-building-scriptingType phenix.python to start up Python with the Phenix environment all set up:
phenix.pythonSet up the high level objects, so we can start our task:
from iotbx.data_manager import DataManager # Load in the DataManager
dm = DataManager() # Initialize the DataManager and call it dm
dm.set_overwrite(True) # Overwrite files with the same name
mmm = dm.get_map_model_manager( # getting a map_model_manager
model_file="short_model_main_chain_box.pdb", # model file
map_files="short_model_box.ccp4") # map file
Fitting a loop
We can fill in the resolution and experiment type and get a model_building object:
mmm.set_resolution(3) # resolution is 3 A
mmm.set_experiment_type('cryo_em') # it is a cryo-EM map
build = mmm.model_building() # get a model-building object
Let’s remove some residues in this model to create a model that needs a loop (we don’t have to do this, but it makes this example clearer):
new_model = build.model().apply_selection_string( # apply a selection
' not (chain A and resseq 516:519)' ) # remove residues 516-519 chain A
build.set_model(new_model) # replace existing model with new_model
You can read about how to specify selections in the Phenix selection syntax documentation. The apply_selection_string method is part of a model object and it applies the selection you specify and returns a new model based on that selection.
Let’s write out the working model:
dm.write_model_file(build.model(), 'model_without_loop.pdb') # new model
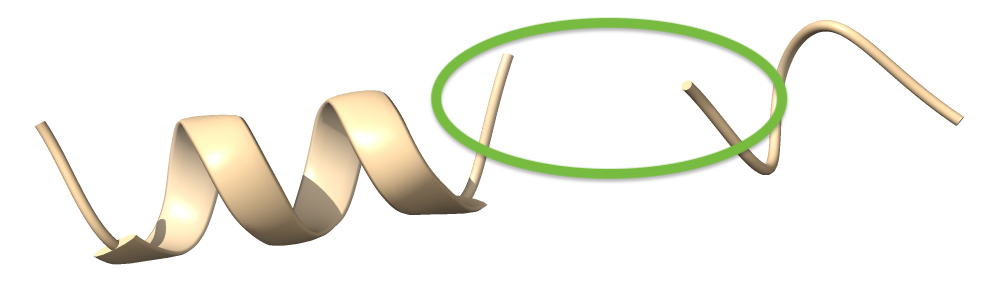

We can choose which methods to use in loop fitting. Let’s use resolve loop fitting:
build.set_defaults(fit_loop_methods=['resolve']) #choose just resolve loop fitting
And let’s fit a loop. If were are already residues where the loop is to go (between residues 515 and 520) they would be removed before the loop is built:
fit_loop_model=build.fit_loop( # fit a loop
chain_id="A", # chain ID
loop_sequence = 'RVLD', # Sequence of residues in loop
last_resno_before_loop=515, # Residue number just before loop
first_resno_after_loop=520,) # Residue number just after loop
Let’s insert the loop:
build.insert_fitted_loop() # insert the loop
Finally let’s write out the new model:
dm.write_model_file(build.model(), 'fitted_loop_model.pdb') # new model